If `pmp_thr` is the only rxd mechanism producing a membrane flux for calcium then simply recording the calcium current (as you have `ca_cur`) is sufficient.
Unfortunately if this is not the case the contribution of the current resulting from a particular MulticompartmentReaction is not directly accessible, but could be obtained using additional rxd.State. For example;
Code: Select all
from neuron import rxd, h
from neuron.units import mV, mM, nM, ms
from matplotlib import pyplot
h.load_file('stdrun.hoc')
soma = h.Section(name="soma")
cyt = rxd.Region([soma], name="cyt", nrn_region="i")
mem = rxd.Region([soma], name="mem", geometry=rxd.membrane)
ecs = rxd.Region([soma], name="ecs", nrn_region="o")
ca = rxd.Species([cyt,ecs], name="ca", charge=2, initial=lambda nd: 1 * mM if nd.region == ecs else 50 * nM)
pump = rxd.Species([cyt,mem], name="pump", initial=1 * mM)
pump_ca = rxd.Species([mem], name="pump_ca", initial=0* mM)
rec_in = rxd.State([cyt], name="rec_in", initial=0)
rec_out = rxd.State([cyt], name="rec_out", initial=0)
k1_pmp, k2_pmp, k3_pmp, k4_pmp = 1e8, 1e4, 1e4, 1e3
pmp_in = rxd.MultiCompartmentReaction(ca[cyt] + pump[mem], pump_ca[mem] + rec_in[cyt],
k1_pmp * ca[cyt] * pump[mem],
k2_pmp * pump_ca[mem],
mass_action=False,
membrane_flux=True,
membrane=mem)
pmp_out =rxd.MultiCompartmentReaction(pump_ca[mem] + rec_out[cyt], ca[ecs] + pump[mem],
k3_pmp * pump_ca[mem],
k4_pmp * ca[ecs] * pump[mem],
mass_action=False,
membrane_flux=True,
membrane=mem)
t_vec = h.Vector().record(h._ref_t)
ca_con = h.Vector().record(soma(0.5)._ref_cai)
ca_cur = h.Vector().record(soma(0.5)._ref_ica)
pup_curi = h.Vector().record(rec_in.nodes(soma)._ref_value)
pup_curo = h.Vector().record(rec_out.nodes(soma)._ref_value)
h.finitialize(-70 * mV)
h.continuerun(100 * ms)
# the flux (in mM/ms) is the derivative wrt time of the record state
flux_in = (pup_curi.as_numpy()[:-1] - pup_curi.as_numpy()[1:])/h.dt
flux_out = (pup_curo.as_numpy()[:-1] - pup_curo.as_numpy()[1:])/h.dt
# the change in cytosolic calcium due to the pump in (mM/ms)
pump_flux = flux_in - flux_out
# as the pump is the only rxd mechanism producing a membrane flux of calcium
# the pump flux can also be calculate by recording the calcium current
pump_flux2 = -0.5e4*ca_cur * soma(0.5).area() / soma(0.5).volume() / h.FARADAY
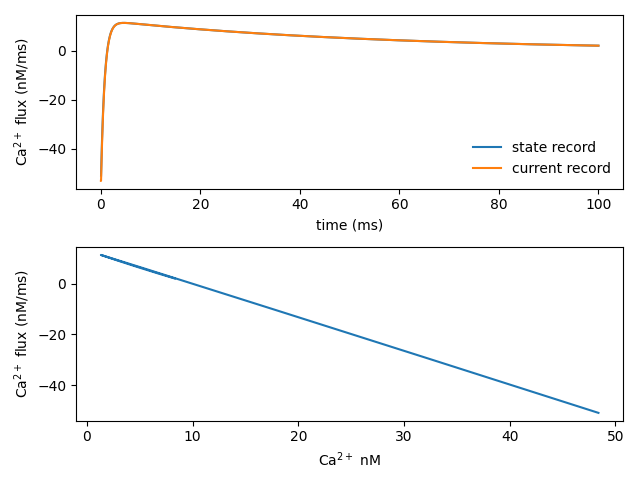
pyplot.figure()
pyplot.subplot(2,1,1)
pyplot.plot(t_vec.as_numpy()[1:], pump_flux / nM / ms, label="state record")
pyplot.plot(t_vec, pump_flux2 / nM / ms, label="current record")
pyplot.legend(frameon=False)
pyplot.xlabel("time (ms)")
pyplot.ylabel("Ca$^{2+}$ flux (nM/ms)")
pyplot.subplot(2,1,2)
pyplot.plot(ca_con.as_numpy()[1:]/nM, pump_flux / nM / ms)
pyplot.xlabel("Ca$^{2+}$ nM")
pyplot.ylabel("Ca$^{2+}$ flux (nM/ms)")
pyplot.tight_layout()
pyplot.show()
Notice the addition of `mass_action=False` to ensure the additional states do not change the dynamics of the pump. Also I think both the `pmp_thr` and `pump_out` should both produce a current, so have set `membrane_flux=True` in each.
The latest documentation is on github;
https://neuronsimulator.github.io/nrn.

