MechanismStandard (Parameter Control)¶ ↑
-
class
MechanismStandard¶ ↑ Syntax:
ms = h.MechanismStandard(name_str) ms = h.MechanismStandard(name_str, vartype)
- Description:
In Python, consider the use of ‘sec.psection()’ which encapsulates MechanismType and MechanismStandard so as to return a dictionary.
With no vartype or vartype = 1, this provides storage for parameter values of a membrane mechanism or point process. This class is useful in maintaining a default set of parameters and can be used to specify values for a set of sections.
name_str is a density mechanism such as
hhor a point process such asVClamp. AMechanismStandardinstance, when created, contains default values for all parameters associated with the mechanism.In combination with the
MechanismTypeclass it is possible to create generic graphical interface widgets that are independent of the particular mechanism and parameter names.If vartype = 1, 2, or 3, the storage is for PARAMETER, ASSIGNED, or STATE variables respectively. If vartype = 0, the storage is for all three types.
If vartype = -1, the count and names (and array size) of the GLOBAL variables are accessible, but any other method will generate an error message.
Example:
from neuron import h, gui ms1 = h.MechanismStandard('hh') ms2 = h.MechanismStandard('AlphaSynapse') ms2.set('gmax', 0.3) ms1.panel() ms2.panel() ms1 = h.MechanismStandard("hh") ms2 = h.MechanismStandard("AlphaSynapse") ms2.set("gmax", .3) ms1.panel() ms2.panel()
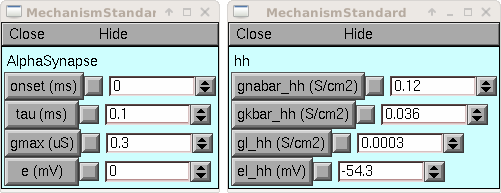
Example:
The following example prints all the names associated with POINT_PROCESS and SUFFIX mechanisms.
from neuron import h, gui soma = h.Section() def pname(msname): s = h.ref('') for i in range(-1, 4): ms = h.MechanismStandard(msname, i) print('\n{} vartype={}'.format(msname, i)) for j in range(ms.count()): k = ms.name(s, j) print('%-5d %-20s size=%d' % (j, s[0], k)) def ptype(): msname = h.ref('') for i in range(2): mt = h.MechanismType(i) for j in range(mt.count()): mt.select(j) mt.selected(msname) print('\n\n{} mechanismtype={}'.format(msname[0], j)) pname(msname[0]) ptype()
Example:
The following example provides a function
get_mech_globalsthat returns a list of all of a mechanism’s global (or per-thread-global) variables. As running the code shows, there are six such variables (all per-thread-global) for thehhmechanism. These are used to temporarily share limiting values and time constant information between functions in the NMODL file; their per-thread-global nature means that the memory is reused for subsequent locations within a given thread, but that different threads do not interfere with each other.from neuron import h def get_mech_globals(mechname): ms = h.MechanismStandard(mechname, -1) name = h.ref('') mech_globals = [] for j in range(ms.count()): ms.name(name, j) mech_globals.append(name[0]) return mech_globals print(get_mech_globals('hh'))
See also
-
MechanismStandard.panel()¶ ↑ - Syntax:
ms.panel() ms.panel("string")
- Description:
Popup a panel of parameters for this mechanism. It’s a good idea to set the default values before generating the panel.
With no argument the first item in the panel will be the name of the mechanism. Otherwise the string is used as the first item label.
See also
-
MechanismStandard.action()¶ ↑ - Syntax:
ms.action(py_callback)
- Description:
- py_callback is executed when any variable is changed in the panel.
The callback is sent three parameters; in order: the MechanismStandard object,
the index of the changed item in the object, and a third argument indicating
position in an array (or 0 if the parameter is not an array; this is the usual
case). The value is in h.hoc_ac_ and this value may also be read via
nameref = h.ref(""); ms.name(nameref, i); value = ms.get(nameref[0], j)
Example:
from neuron import h, gui soma = h.Section(name='soma') axon = h.Section(name='axon') dend = [h.Section(name='dend[%d]' % i) for i in range(3)] axon.insert('hh') for sec in dend: sec.insert('pas') h.xpanel("Updated when MechanismStandard is changed") for i, sec in enumerate(dend): h.xvalue("dend[%d](0.5).pas.g" % i, sec(0.5).pas._ref_g) h.xpanel() def change_pas(ms, i, j): for sec in h.allsec(): if sec.has_membrane('pas'): ms.out() ms = h.MechanismStandard('pas') ms.action(change_pas) ms.panel()
Note
Support for Python callbacks for this method was added in NEURON 7.5.
-
MechanismStandard._in()¶ ↑ - Syntax:
ms._in(sec=section) ms._in(x, sec=section) ms._in(pointprocess) ms._in(mechanismstandard)
- Description:
copies parameter values into this mechanism standard from …
ms._in(sec=section)- the mechanism located in first segment of
section ms._in(x, sec=section)- the mechanism located in the segment
section(x). (Note that x=0 and 1 are considered to lie in the 0+ and 1- segments respectively. ms._in(pointprocess)- the point process object
ms._in(mechanismstandard)- another mechanism standard
If the source is not the same type as the standard then nothing happens.
Example:
from neuron import h s = h.Section(name='soma') s.insert('hh') s(.5).hh.gnabar = 0.5 ms = h.MechanismStandard('hh') ms.set("gnabar_hh", 0.3) print(ms.get("gnabar_hh")) ms._in(sec=s) print(ms.get("gnabar_hh"))
Note
This is the same as the HOC method
ms.in, however the name had to be changed for Python due toinbeing a keyword in Python.Note
Python support for this method was added in NEURON 7.5.
-
MechanismStandard.out()¶ ↑ - Syntax:
ms.out(sec=section) ms.out(x, sec=section) ms.out(pointprocess) ms.out(mechanismstandard)
- Description:
copies parameter values from this mechanism standard to …
ms.out(sec=section)- the mechanism located in
section(all segments). ms.out(x, sec=section)- the mechanism located in
sectionin the segment containing x.(Note that x=0 and 1 are considered to lie in the 0+ and 1- segments respectively) ms.out(pointprocess)- the point process argument
ms.out(mechanismstandard)- another mechanism standard
If the target is not the same type as the standard then nothing happens.
-
MechanismStandard.set()¶ ↑ - Syntax:
ms.set('varname', val [, arrayindex])
- Description:
sets the parameter in the standard to val. If the variable is an array, then the optional index can be specified.
varnamefollows the HOC form convention ofname_mech; e.g.gnabar_hh.See
MechanismStandard.out()for an example.
-
MechanismStandard.get()¶ ↑ - Syntax:
val = ms.get('varname' [, arrayindex])
- Description:
returns the value of the parameter. If the variable is actually a POINTER and it is nil, then return -1e300.
varnamefollows the HOC form convention ofname_mech; e.g.gnabar_hh.See
MechanismStandard._in()for an example.
-
MechanismStandard.save()¶ ↑ - Syntax:
ms.save('name')
- Description:
- For saving the state of a MechanismStandard to a session file. The name will be the objectvar that the instance gets assigned to when the session file is read. See pointman.hoc for an example of usage.
-
MechanismStandard.count()¶ ↑ - Syntax:
cnt = ms.count()
- Description:
- Returns the number of parameter names of the mechanism represented by the MechanismStandard.
-
MechanismStandard.name()¶ ↑ - Syntax:
ms.name(strref) size = ms.name(strref, i)
- Description:
The single arg form assigns the name of the mechanism to the strref variable.
When the i parameter is present (i ranges from 0 to ms.count()-1) the strref parameter gets assigned the ith name of the mechanism represented by the MechanismStandard. In addition the return value is the array size of that parameter (1 for a scalar).
Example:
from neuron import h, gui ms = h.MechanismStandard('hh') name_strref = h.ref('') # read the name of the mechanism ms.name(name_strref) print(name_strref[0]) # displays: hh