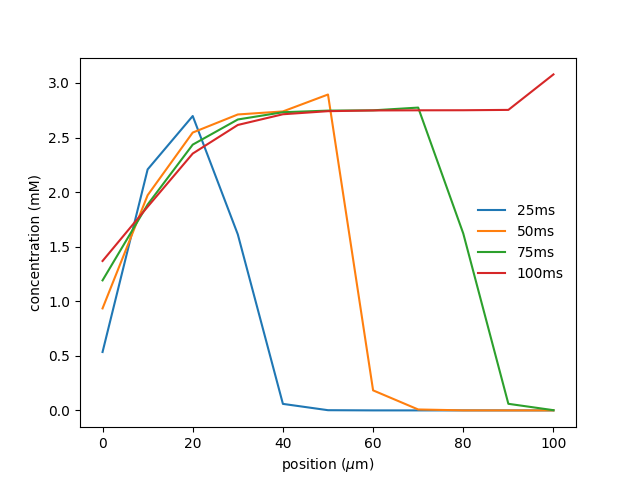
I am currently working on a Python rxd simulation where we are simulating multiple mitochondria being transported within an axon, while also keeping track of their molecular outputs (e.g. ATP production and calcium buffering) in both the spatial (1D) and temporal dimension.
I have two questions I need help in.
Question 1: My first question is how I should best implement such a model. Essentially I have one large region (axon), with several moving compartments within it (mitochondria), where it is important to keep track of not only their internal parameters individually, but also their effect on the axon is localized to their position.
My current implementation is to divide the axon Region into Sections of fixed lengths. Then I had defined internal molecular Species (e.g. mitochondrial ATP, Ca) for each individual mitochondria using Species(....,initial = function(node),...),using the function to define the Species as having a value at the mitochondrial position node and 0 everywhere else - essentially my way of saying "all the stuff is here". Then during my simulation, if it detects that a mitochondria's location has resulted in it changing nodes, it shifts the Species value within the array where appropriate.
As for the Reaction Rates (e.g. ATP and Ca transport), I had to define Rates for each individual mitochondria (because they are often dependent on the mitochondria's internal Species concentration). I also used an rxd.Parameter which marks the active node as 1 (where the mitochondria is) and the rest 0, so I can multiply that with the Rate and thus make it only work at that specific node.
While this implementation seems to work fine (for the most part, my 2nd question regards a more specific issue regarding setup), my gut instinct tells me that there could be a better way to do this.
Question 2: My second, more specific question is about setting up the Rates.
Our mitochondria molecular rates are based on papers and we had recently discovered a new paper that showed issues in the simplified equations that we had been using thus far.
However, implementing the new equations resulted in a rather bulky reaction rates. For example:
Code: Select all
# Calcium Reaction Rates, From MK (Bertram), verified,unit: μM/ms
calcrate = []
for i in range(Mito_num):
ratedict = {}
ratedict["u"] = cac/0.2
ratedict["G(u)"] = ratedict["u"]*((1+0.52*ratedict["u"])**2.8)*((1+0.01*ratedict["u"])**3)/(110+((1+0.52*ratedict["u"])**2.8)*(1+0.01*ratedict["u"])**4)
ratedict["w"] = mitolist[i]["PSIm"]/91
ratedict["J_uni_MK"] = ratedict["G(u)"]*0.11*6.73*(ratedict["w"]-1)/(1-rxdmath.exp(-6.73*ratedict["w"]-1))
#new equation from Siqueria2013
ratedict["J_NCX_MK"] = 0.06*(mitolist[i]["cam"]/cac)*rxdmath.exp(0.018*(mitolist[i]["PSIm"]-91))/((1.94**3)*(1+0.375/mitolist[i]["cam"]))
ratedict["Uni_in"] = rxd.Rate(mitolist[i]["cam"], g_ca*ratedict["J_uni_MK"]*mitolist[i]["node_act"])
ratedict["NCX_out"] = rxd.Rate(mitolist[i]["cam"], -g_ca*ratedict["J_NCX_MK"]*mitolist[i]["node_act"])
calcrate.append(ratedict)Based on Spyder profiler, the problem appears to arise from the following check in rate.py, line 63.
Code: Select all
self._voltage_dependent = False if not hasattr(rate,'_voltage_dependent') else rate._voltage_dependent
Given the bulk of my equation (probably not helped by having individual rates for individual mitochondria), it results in this check being made a huge number of times. If I change the code in rate.py to:
Code: Select all
self._voltage_dependent = False
Thank you for your help and ask me if you need more information!